Text Mining and Natural Language Processing on Health Forums
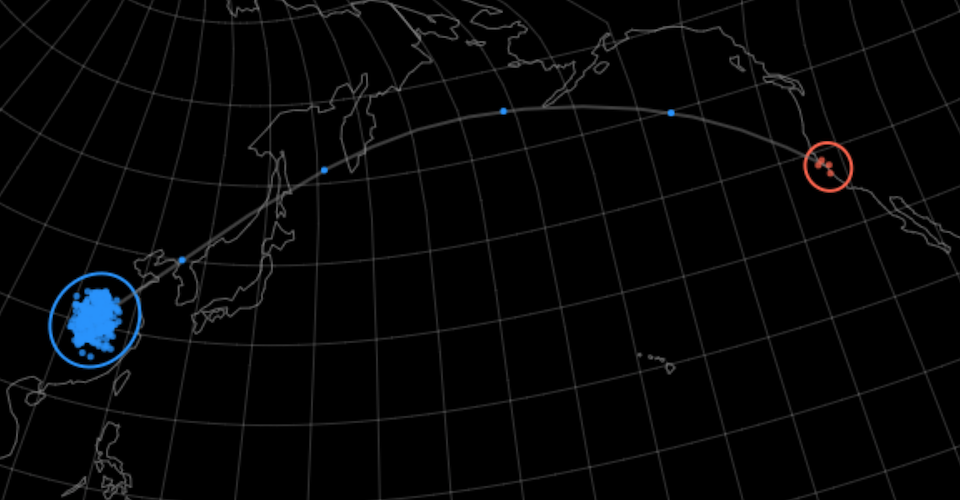
As part of the Insight Health Data Science Fellowship, I just got to spend the last 3 weeks working on a pretty fun project applying natural language processing to medical health forums. The goal is to curate health forums so that people can get instant advice on sexual health problems and then see the most relevant forum posts on those issues.
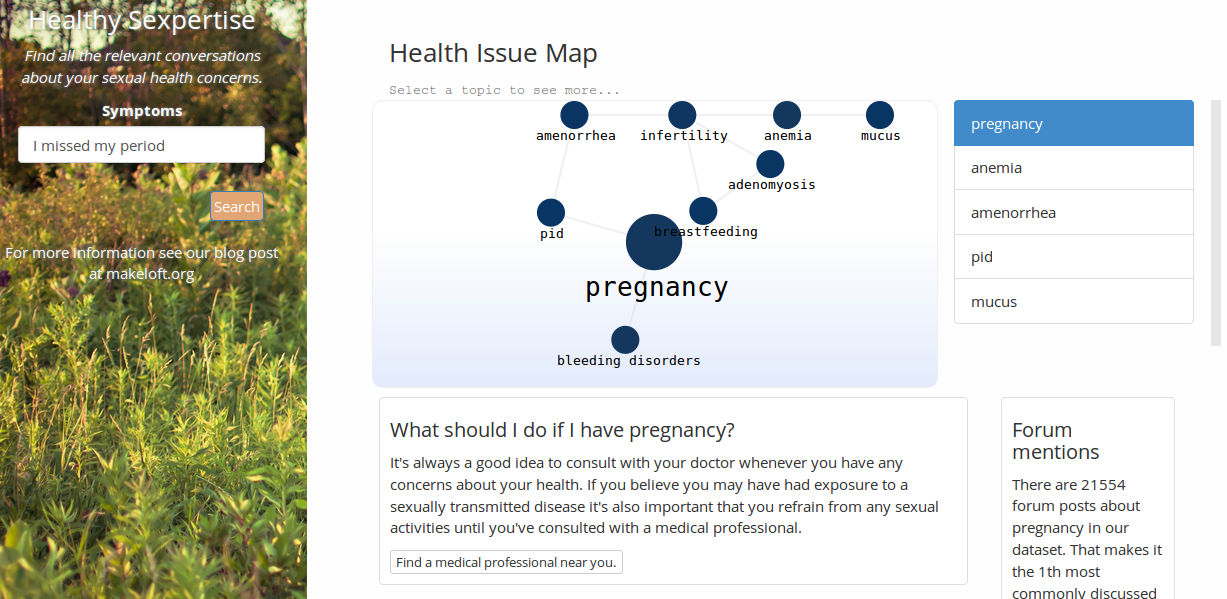
The web app is up and running. To try it out, hop on over to sexpertise.makeloft.org (maybe use an incognito tab if you're skiddish about your search history). I'll eventually do another post all about the AWS backend API and the D3 frontend stuff, but for this post I want to focus in just on the text mining and analysis that went into building the condition suggester. To jump straight to the code you can check out my Github repo.
Health Forum Datasets
At least 4 out of 5 internet users ask health related questions online and google says 1 in 20 searches are for medical info. Clearly, online information is an integral part of the healthcare pipeline in this day and age. Providing better information to users up front can change their whole medical experience downstream.
Nowhere is this more important than in sexual health information because social taboos can prevent people from feeling comfortable seeking medical help. As many as 63% of people who go online for medical advice say they turn to the internet to talk about sensitive issues like sex and STDs. Fortunately, the internet is helping to alleviate some of these constraints.
To get advice on these issues many users turn to online "Ask a Doctor" forums found on WebMD, DoctorsLounge or eHealthForum. All of these sites offer free anonymous questions with medical professionals. On these sites, typically an "Asker" poses a question in a public forum, which can then be answered by a doctor.
As an example let's look at the following interaction between an asker and a doctor, which can be found at this doctorslounge.com post.

The asker normally talks about symptoms that he or she is experiencing (blue), while the doctor responds with suggestions and, importantly, mentions different likely conditions (green). Unfortunately, it can take as much as a week for a doctor to respond, and this can cause the asker to wait unnecessarily rather than seeking medical help at a clinic.

It would be great to be able to mine the text so that we could map those symptoms to conditions automatically. At the same time, it's clear that there's a lot of extra information in the posts. Without any further analysis, it's basically impossible to tell what's important in these messages and how to extract meaning from them. But fortunately, there is a process for working with textual data known as NLP that will help us make sense of this.
Scraping forum data
In order to analyze these forums I needed to download many forum conversations from multiple websites. To do this I used BeautifulSoup to crawl the webforus and store the text from the asker, the doctor, and any other respoonders. I ended up getting data from WebMD, eHealthForum, DoctorsLounge, and ScarletTeen sexual health forums. Others that I could add would be reddit and stackexchange as well as medhelp.
I downloaded 150,000 posts in 60,000 threads and about half of them had a response from a medical professional (either a doctor or a registered nurse). Identifying medical professionals was always specific to each forum, and the best one can do is get a sense of the heuristics each site uses to identify their "official" responses. To view the scrapers that I used to download the data check out my insight-health-scraper repo on GitHub.
Filtering on writing style
One of the first things I noticed when perusing my dataset was the wide disparity in the quality of Asker questions and their writing style. I also noticed that the doctors tended to sound a lot smarter than the askers as you might expect. This led me to wonder whether I could detect the quality of the writing style. To do this I calculated the Flesch-Kincaid Grade Level of each post in my corpus using mmauter's readability package for python. The Flesh-Kinkaid readability tests are a system to estimate the reading ease of paragraphs. The Grade Level tries to map someone's writing onto the equivalent educational Grade Level in the American school system. I applied it to my corpus and was able to get a nice plot of the frequency of reading levels for both the asker and doctor texts.
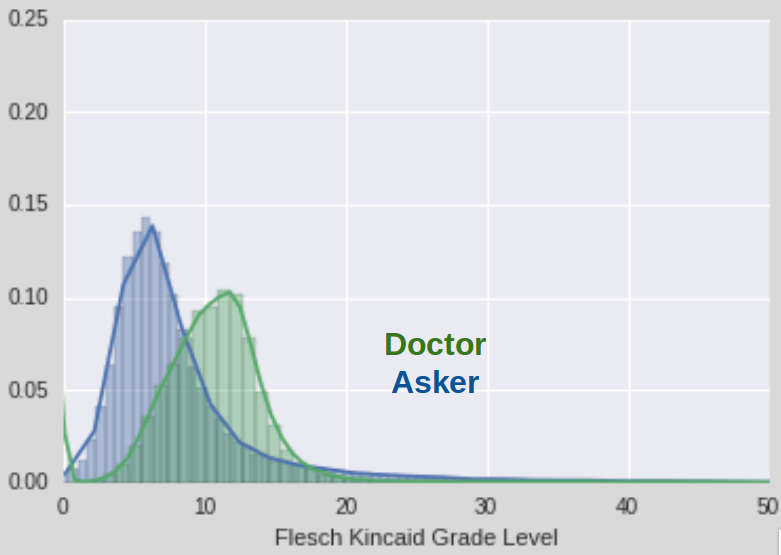
As you can see, using this metric, doctor texts score significantly higher than asker texts. Flesch-Kincaid mainly takes into account the number of syllables in words and the number of words in sentences. Ernest Hemingway would probably disagree that those are good metrics for the quality of writing, but it's a fair proxy short of any other information about the text. For example, one asker posed the following thought provoking health question:
Should I have sex with some random guy?
This sentence actually scores negative on the Flesch-Kinkaid grade level. To me it seems like it should score negative on pretty much every level.
At any rate, the Flesch-Kincaid metric gave me a method to establish which asker questions were just unfit to be displayed. At the end of the project, this helped in filtering the responses that I showed to end users. However, I still left that data in during the modelling part.
Finding Conditions in Doctor Responses
After the dataset is downloaded and tidied, you have to ask what you are going to do with the dataset. At the moment, the whole dataset just consists of pairs of unstructured text, one for asker and one for doctor. You could try to map from asker text to doctor text (see below for that problem), but first it would be better to get a simpler set of labels to map to. In the end, what I really wanted was just a list of all the recognizable conditions that the doctor mentions.
So to turn this into a more cleanly labelled dataset, I needed to generate a list of condition labels directly from the doctor text. This turns out to be not a trivial task with lots of ways to tweak the final result. I can't offer a perfect answer, but I can offer the solution I used to detect conditions in doctor text.
Generating condition synonym list from CDC and Mayo clinic websites
To get recognizable conditions to label, I needed to build a list of text strings that could be construed to represent "diseases". Fortunately, both the CDC and Mayo Clinic have lists of diseases and conditions which were easy enough to scrape using my insight-health-scraper package.
After I had the raw list, I realized that there were lots of conditions that were pretty similar to each other. I decided I needed to collapse this list down to a shorter list of truly unique entries for two reasons: 1) I didn't want an overwhelming array of conditions to display at the end, and 2) I wanted to enhance the predictive ability of my model by increasing the number of occurrences of each condition I was trying to model. To do this I used a technique called fuzzy matching to see which strings were more than 90ish% similar to each other.

I relied on a cute python package called fuzzywuzzy, which worked great for identifying close partial matches of strings. Specifically, I could just use the extractOne() method to find the closest match out of the diseases I've already seen. If the match was better than 90%, just add it as a synonym to the original disease, creating a way to map all of the similar strings to one term.
The many ways to match a string
Once I had the disease synonyms, I needed to look into the doctor text to see when they were mentioned. There were a number of different ways to do this, and I explored several permutations of the following techniques.
Multiple "synonyms"
I already had multiple different terms that mapped to the same disease so I had a long list of somewhat similar terms that could be compared against (e.g. 'ehrlichiosis' and 'human ehrlichiosis' both map to the same disease).
Spelling correction
Another cool package called PyEnchant can make spelling corrections. It is slow so I didn't use it on the whole dataset, but it could offer another way to correct spelling errors for diseases.
Fuzzy matching
As I mentioned above, fuzzy matching allows similar strings that are not exact matches to be compared. Again, this technique is much slower than a simple string comparison, but I decided to use it anyway in this case to make sure I didn't miss any similar text.
Marking negation
Finally, it's possible that a mention doesn't necessarily mean the text is relevant. In particular, a doctor could be saying that she doesn't think you have a condition (e.g. "You probably don't have an STD"). Using negation marking in nltk.util allows you to mark every word in a sentence that comes after a negation (i.e. not, no, etc). I stored the results of this, but it only reduced the number of identified conditions by 1% so I didn't bother using it for simplicity of explanation. Nevertheless, it can be a good idea if you're trying this on a different dataset.
Disease labelling results
I searched for 200 diseases across the 30,000 doctor texts. Of those, only 80 occurred more than 100 times so I decided to just focus on those. The following diagram shows some of the most and least frequently mentioned diseases./p> 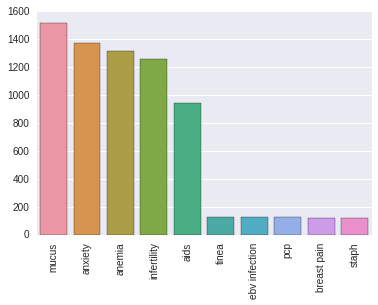
I also repeated this with the asker comments to see which conditions were mentioned by them. In addition I queried the top disease-related wikipedia articles to see how many times they were viewed and I compiled some CDC data to see which diseases occurred the most. Combining these, it was interesting to see what the most commonly mentioned issues were from these disparate datasets

Clearly different things are on different people's minds.
Predicting Conditions from Questions
With a labelled dataset, I could generate a model that predicted conditions using input text. However, the asker text was still just an unprocessed string, which isn't really easy to model without to measure the occurrence of important words. Unfortunately, there is no obvious way to extract the most important words from the text a priori. Instead, I had to use some natural language processing techniques to extract the features of the text into a numerical form that could be fit with a model.
Term Frequency Inverse Document Frequency (TFIDF)
The most traditional way to convert text into a vector of features is called the "bag of words" model. In this model, you lose all information about the ordering of the words and you basically just end up with counts of occurrences of each word. I used a slight adaptation of this called TFIDF, where you normalize the word counts by the frequency of those words across all the documents in your corpus. This has the effect of making it so that your representation has values much larger than 1 whenever a word is over-represented compared to the average document.
You can find a lot of information about TFIDF on the web. It is the standard and most obvious way to deal with this problem. Just to briefly make it clear what we got out of this. Basically after converting your text into TFIDF form, you'll have a vector of numbers where each element in the vector represents how over-represented a certain word is in that text. One consequence is that two identical documents will have identical vector representations. Similarly documents with similar word frequencies will have simimilar vector representations.
Stop Words, Stemming, and Ngrams
There are three additional techniques that extend the usefulness of TFIDF. First, removing stop words filters out the most common English words that don't really convey meaning. Words like 'I', 'in' and 'the' get dropped so they aren't wasting space in your vectorized representation. Second, stemming drops inferred suffixes off of words. Thus, 'running,' 'runs,' and 'run' will all map to 'run,' which further reduces your feature space. Finally, generating ngrams creates new words out of continuous segments of words. So for example, without ngrams you would have a count of 'bus' and a count of 'driver' in your document, but you wouldn't know if the term 'driver' was ever associated with 'bus.' If you use a bigram (or 2-gram) then you would also have the count of 'bus driver' in your text. This can give you more meaning if t turns out to be an important feature.
Restricting feature frequency
I also filtered out words and phrases that were too common and too rare. To do this I clipped out the least frequent 1% and the most frequent 40% of words. This was something where I really wanted to explore the effect on my model systematically, but due to time constraints I had to move on after finding a "good enough" restricting condition.
Experiments with Doc2Vec
In addition to the classic TF-IDF, I also experimented with the state of the art gensim Doc2Vec for extracting features from my text. Doc2Vec (and its basis Word2Vec) is a complicated algorithm that many other people have attempted to explain better than me. In a nutshell, it attempts to look at the context.
One great thing about it is that it maps different words into a "meaning" space. This means that you can map synonymous words to the same feature space with lower dimensionality. As such different words can be have their nearby regions inspected and you can see which words are the most similar in meaning. Here I display 3 different Doc2Vec implementations, and which words they find closest in meaning to ovulation.

In truth, Doc2Vec is a family of algorithms, each with its own pluses and minuses. As you can see the one on the left looks to be the most informative. That one uses the skip-gram model, which you can look into if you are interested in the details. All in all, I'm impressed with Doc2Vec, and with a little more fiddling I'm sure I could change my implementation to include this more elegant vectorization procedure. However, the current Doc2Vec implementations do not enable you to directly map a text to a unique vector representation. Instead, they infer a "good enough" representation with some random noise, which is not guaranteed to be the same every time you query. I didn't have time to fix this issue so I left the TFIDF in production.
Results of the vectorization
To get a better idea of what comes out of the vectorization process take a look at this text from the example above.

As you can see, many of the words have been truncated by the stemming. Also, even though there are many possible ngrams in this example, none of them are in the final vectorization. That just means that they were either too frequent or not frequent enough to make the cut. All in all, a lot of information has been lost, but we don't know if we've kept enough until we've tried to make a model.
Modelling sparse data with multiple labels
For this model, I didn't have a lot of data, but I had a lot I had to predict. In the end, I was trying to predict 80 diseases, and many of those diseases didn't even have many more than 100 examples in the dataset. That leaves a lot of room to get things wrong.
For example, using the F1 score I only had actual predictive power over a handful of conditions. This was universally true with any model that I used (random forests, SVM, logistic regression, and naive bayes) which is a good indication that I just didn't have enough samples to get a predictive model.

Fortunately, after a little while I realized that my problem was actually way easier than trying to predict the diseases. Really, all I needed was to identify the 10-or-so most probable diseases associated with a given disease. Doing that with some fidelity isn't anywhere near as hard, and I found that I could be pretty successful even with my small dataset. The resulting conditions definitely passed the gut test and matched with pretty good fidelity to my expectations.
To be more systematic, I used an ROC area under the curve to score my models and found that I could get 0.85 (1 being the best) using a moderately sized random forest. So that was the model that ended up going into production.

Viewing the most predictive words for each condition
One interesting thing that falls out of this modelling is the ability to see which words are most important for predicting a given condition. I built an interactive analysis tool to visualize the words that are best at predicting a given condition. One of the coolest predictions is for appendicitis.

As you can see, two of the most closely related phrases are 'right' and 'abdominal pain.' To me it's absolutely incredible that this ended up working so perfectly. Feel free to explore the analysis tool to find more cool patterns like this.
Bonus: Disease Connectivity Map
As a cool little additional analysis, I also found that I could infer the connections between diseases based on their co-occurrence in doctor responses. Using this connection map I was able to build a little D3 force graph to visualize the connectivity.

You can definitely see clusters amongst some obviously related conditions. Women's health issues like pregnancy, breastfeeding, and back pain; flu, sore throat, and cold; and all the stds clump together. Some things are less explainable. If you want to explore first hand you can go to the analysis page. Have fun and be safe.
 will stedden
will stedden